US researchers use IBM supercomputer, find 77 molecules that deter COVID-19 infection
By Lim Jeong-yeoPublished : March 12, 2020 - 17:00
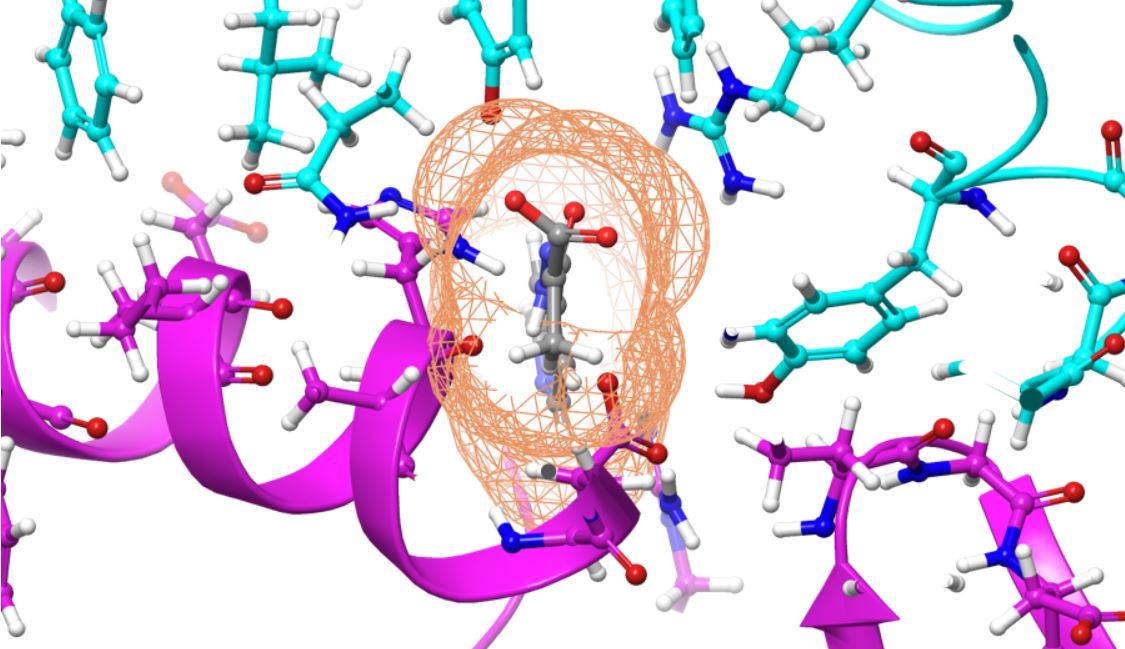
US researchers from the University of Tennessee have identified 77 molecules that show potential to impair COVID-19’s ability to infect host cells, IBM said Thursday. They made the discovery with the help of simulations using supercomputer Summit.
Viruses like COVID-19 infect cells by binding to them and using “spikes” to inject their genetic material into the host cells.
To understand a new microorganism, researchers grow it and test its reaction to new compounds.
The process is a very slow one, but with computers they can perform digital simulations to narrow down variables.
Using Summit, researchers were able to simulate 8,000 compounds in a matter of days and identified 77 small-molecule compounds, such as medications and natural compounds, that showed potential to impair COVID-19’s ability to invade host cells, said Dave Turek, vice president of technical computing at IBM Cognitive Systems.
“Summit was needed to rapidly get the simulation results we needed. It took us a day or two whereas it would have taken months on a normal computer,” said Jeremy Smith, Governor’s Chair at the University of Tennessee, director of the Center for Molecular Biophysics and principal researcher in the study.
“Our results don’t mean that we have found a cure or treatment for COVID-19. We are very hopeful, though, that our computational findings will both inform future studies and provide a framework that experimentalists will use to further investigate these compounds,” Smith said.
By Lim Jeong-yeo (kaylalim@heraldcorp.com)
Viruses like COVID-19 infect cells by binding to them and using “spikes” to inject their genetic material into the host cells.
To understand a new microorganism, researchers grow it and test its reaction to new compounds.
The process is a very slow one, but with computers they can perform digital simulations to narrow down variables.
Using Summit, researchers were able to simulate 8,000 compounds in a matter of days and identified 77 small-molecule compounds, such as medications and natural compounds, that showed potential to impair COVID-19’s ability to invade host cells, said Dave Turek, vice president of technical computing at IBM Cognitive Systems.
“Summit was needed to rapidly get the simulation results we needed. It took us a day or two whereas it would have taken months on a normal computer,” said Jeremy Smith, Governor’s Chair at the University of Tennessee, director of the Center for Molecular Biophysics and principal researcher in the study.
“Our results don’t mean that we have found a cure or treatment for COVID-19. We are very hopeful, though, that our computational findings will both inform future studies and provide a framework that experimentalists will use to further investigate these compounds,” Smith said.
By Lim Jeong-yeo (kaylalim@heraldcorp.com)


![[Herald Interview] 'Amid aging population, Korea to invite more young professionals from overseas'](http://res.heraldm.com/phpwas/restmb_idxmake.php?idx=644&simg=/content/image/2024/04/24/20240424050844_0.jpg&u=20240424200058)






![[Hello India] Hyundai Motor vows to boost 'clean mobility' in India](http://res.heraldm.com/phpwas/restmb_idxmake.php?idx=644&simg=/content/image/2024/04/25/20240425050672_0.jpg&u=)







